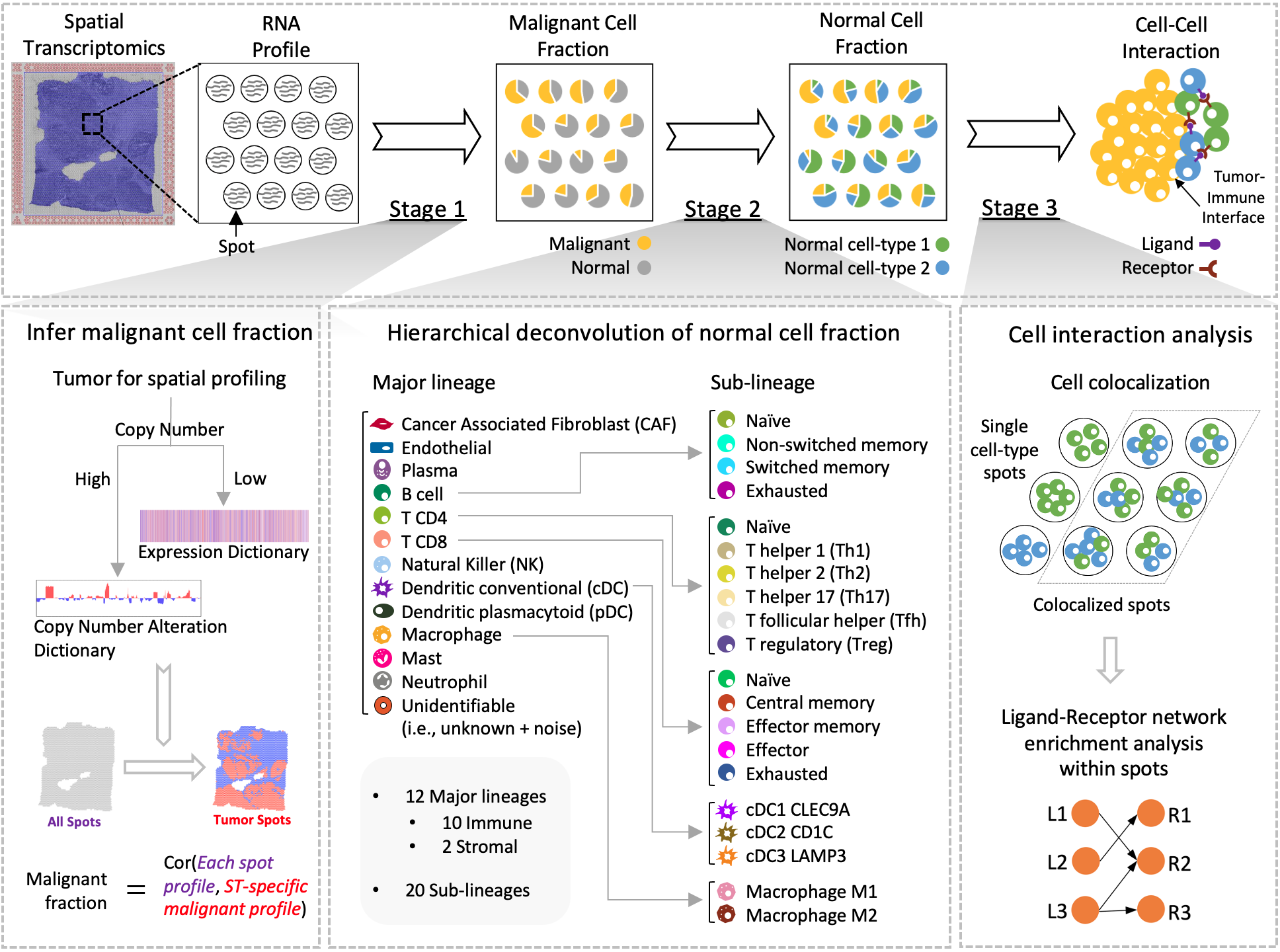
SpaCET is an R package designed for analyzing cancer spatial transcriptomics (ST) datasets to estimate cell lineages and intercellular interactions within the tumor microenvironment. In a nutshell, SpaCET first infers cancer cell abundance by integrating a gene pattern dictionary of common malignancies. Subsequently, SpaCET employs a constrained linear regression model to calibrate local tissue densities and determine stromal and immune cell lineage fractions based on a comprehensive non-malignant cell atlas. Furthermore, SpaCET has the capability to unveil putative cell-cell interactions within the tumor microenvironment, particularly at the tumor-immune interface. Of note, although SpaCET does not require any input cell references for the analysis of tumor ST data, SpaCET can still incorporate a matched scRNA-seq dataset as customized references to conduct cell type deconvolution of any ST dataset. Check the tutorials below for details on how to use the package.
Installation
To install SpaCET, we recommend using devtools:
# install.packages("devtools")
devtools::install_github("data2intelligence/SpaCET")Or user can install SpaCET from the source code. Click here to download it.
# install.packages("remotes")
remotes::install_deps("Path_to_the_source_code", force = TRUE)
# install SpaCET in the R environment.
install.packages("Path_to_the_source_code", repos = NULL, type="source")Dependencies
- R version >= 4.2.0.
- R packages: Matrix, jsonlite, ggplot2, reshape2, scatterpie, patchwork, png, shiny, plotly, DT, MUDAN, factoextra, NbClust, cluster, parallel, pbmcapply, psych, BiRewire, limma, arrow, UCell, RANN, sctransform.
Example
library(SpaCET)
visiumPath <- file.path(system.file(package = "SpaCET"), "extdata/Visium_BC")
SpaCET_obj <- create.SpaCET.object.10X(visiumPath = visiumPath)
SpaCET_obj <- SpaCET.deconvolution(SpaCET_obj, cancerType="BRCA", coreNo=6)
SpaCET_obj@results$deconvolution$propMat[1:13,1:5]Tutorial
SpaCET is applicable for deconvolving spatial transcriptomics data across different platforms and cellular resolutions. In addition, it can compute gene set scores and assess spatial correlations.
Data availability
This Link provides access to the 10 scRNA-seq datasets used to generate SpaCET’s in-house cell-type reference, along with the 8 spatial transcriptomics samples demonstrated in our manuscript.
Contact
For questions, bug reports, or feature requests, please submit an issue. To keep the issue tracker focused and constructive, advertising or promotional content is not permitted.
Citation
Beibei Ru, Jinlin Huang, Yu Zhang, Kenneth Aldape, Peng Jiang. Estimation of cell lineages in tumors from spatial transcriptomics data. Nature Communications 14, 568 (2023). [Full Text]
